PIPELINE · SOIL|WATER|SPECIMEN TO CLAIM
Six stages. One verifiable chain of custody.
From a raw FASTA or DwC-A (Darwin Core Archive) through to a published, data-tethered scientific artifact — every step recorded, every claim re-derivable. The same chain runs under the BioKEA molecular sequencing service, so customer samples land with the same verifiable output as our own.
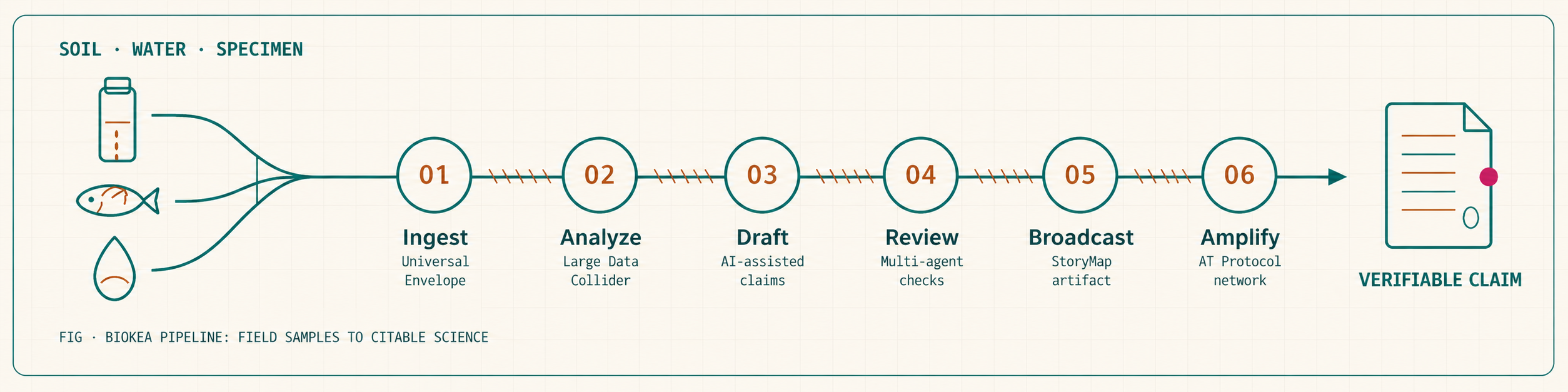
- 01IngestUniversal Envelope
Every input — raw FASTA, DwC-A archive, drafted manuscript — becomes a cryptographically trackable object. Automatic file-type detection and metadata extraction.
- 02AnalyzeLarge Data Collider
The LDC runs image QC, taxonomy reconciliation, and FAIR validation over millions of reads in minutes. Outputs operational taxonomic units and candidate novel lineages.
- 03DraftAI-assisted narrative
The scientist directs; the AI drafts structure and links LDC data directly into the text. Cross-references with external hypotheses in real time.
- 04ReviewMulti-agent panel
AI pre-screens manuscript structure and methodology in hours. Verified human experts evaluate contextual scientific nuance. Weighted, transparent scoring.
- 05BroadcastInteractive StoryMap
The end product is not a dead PDF. It is an explorable digital artifact permanently tethered to its underlying FAIR data package (GBIF, NCBI SRA, Zenodo).
- 06AmplifyATProto / Bluesky
Publishing is the starting line. Seamless AT Protocol integration pushes verifiable scientific artifacts into decentralized social graphs.
BEING BUILT
BioinfoOS
The software layer running on the BioKEA Large Data Collider (LDC). In-house AI-assisted modules cover:
- · Extraction-run QC (Claude Vision over plate images)
- · Taxonomy reconciliation against BOLD, NCBI, and GBIF
- · FAIR (Findable, Accessible, Interoperable, Reusable) package validation — DwC-A, Darwin Core, Zenodo DOI-ready
- · Draft narrative generation tethered to pipeline outputs
- · Operational Taxonomic Unit (OTU) clustering, amplicon denoising, and chimera filtering
Modules ship incrementally; BioinfoOS is in active development and runs on the same LDC hardware used by the molecular sequencing service.
PUBLISHED AT
Agentis
Pipeline outputs publish to Agentis, our forthcoming AI-first open-access platform on the AT Protocol — in early development.
agentis.science →TRUST
Cryptographic provenance, end to end.
Every artifact carries an AT Protocol Decentralized Identifier (DID). Every peer review is a signed, verifiable record. The pipeline doesn't just produce findings — it produces evidence that's re-derivable from raw input to published claim, by anyone, at any time.
Want to plug a sample into this?
We're onboarding sample streams and collaboration partners.
